Beyond the tree: phylogenetic comparative methods for evolutionary inference of phenotypic data
“Nothing in Biology Makes Sense Except in the Light of Evolution” was famously said by Theodosius Dobzhansky. And equivocally, phenotypic variation among
Speakers
Event series
Content navigation
RegisterDescription
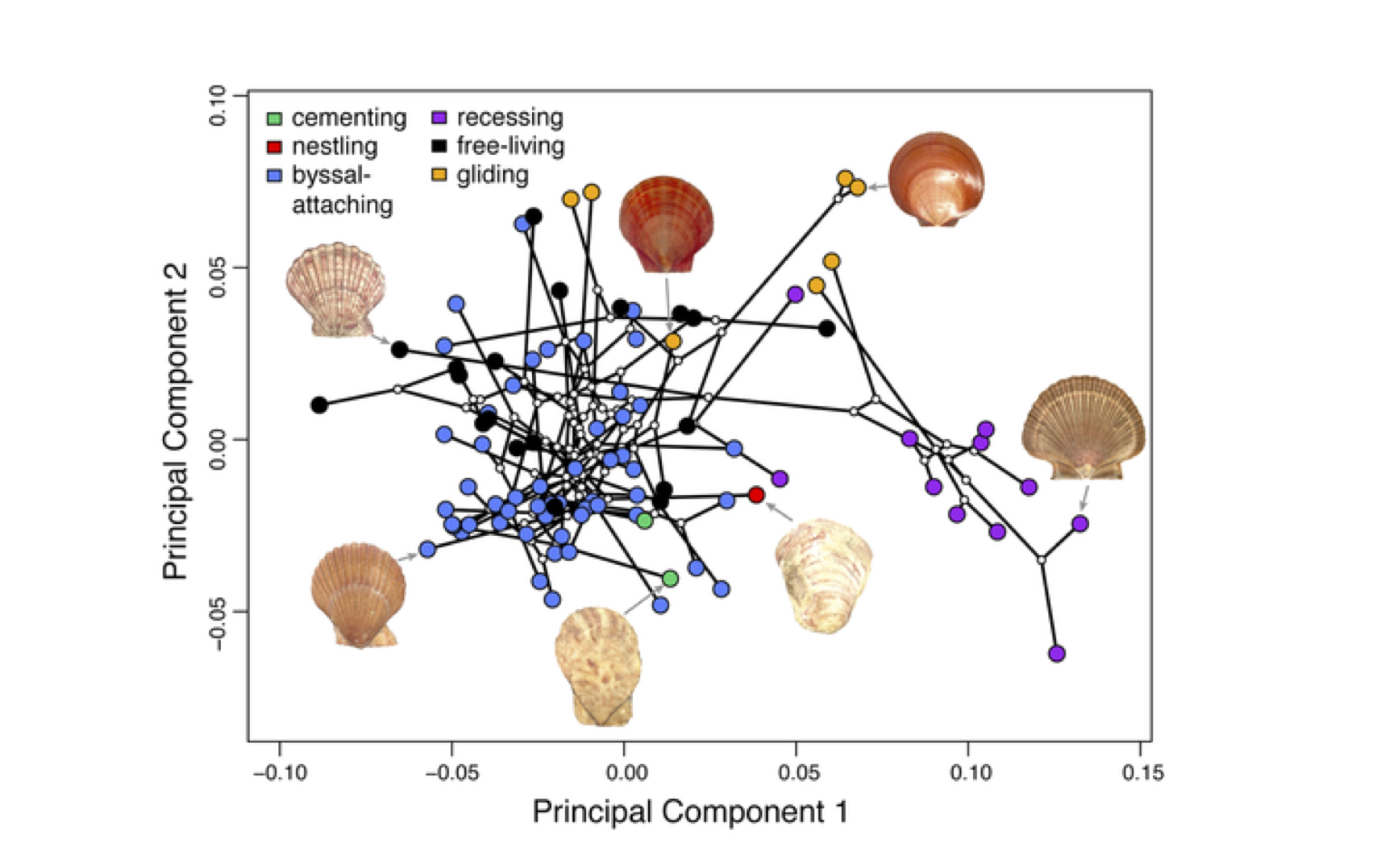
“Nothing in Biology Makes Sense Except in the Light of Evolution” was famously said by Theodosius Dobzhansky. And equivocally, phenotypic variation among species can only be interpreted in light of a phylogenetic framework.
This is a very active field of method development and implementation; journals such as Evolution, American Naturalist and Systematic Biology are built upon this framework. As with all active fields with (often) conflicting or controversial new methods, it can be daunting to navigate without assistance.
Therefore, in this talk “Beyond the tree”, we offer a beginner’s guide, introducing the methods and theory behind mapping phenotypic traits onto phylogenetic trees and statistical analyses of species data (known as phylogenetic comparative methods, PCM).
We will begin by talking about best practices for collecting data for PCM. Then we will use published examples of hypotheses and questions regarding phenotypic trait evolution to demonstrate the current available methods – including reference to implementation with packages in the R statistical framework. Starting with the simplest model of evolution, Brownian Motion (BM), and then progressing to advanced methods that go beyond BM. Finally, we will discuss what the future holds for these approaches.
'Must-have' books on this topic:
- Felsenstein, J. Inferring phylogenies. Vol. 2. Sunderland: Sinauer associates, 2004.
- Garamszegi, L.Z. Modern phylogenetic comparative methods and their application in evolutionary biology. Concepts and Practice. London, UK: Springer 2014.
People from all academic backgrounds and levels are welcome.
We appreciate you taking the time to register (it's free) - it gives us an idea of who’s attending our TEA Talks, and the presenters some background on their audience.
Hosted by the Centre for Biodiversity Analysis, TEA Talks (Techniques in Evolutionary Analysis) are a monthly series of short workshops that introduce a range of current methods and analytical approaches in phylogenetics, bioinformatics and macroevolution.
Location
EEG Seminar Room, Gould Building 116, Daley Rd, ANU
